A team of scientists from the Life Sciences Center of Vilnius University (VU LSC), implementing a project funded by the Research Council of Lithuania, recently published a methodological article in the international journal Febs Open Bio, presenting a method that enables the study of DNA – protein interactions at the level of single molecules using advanced optical fluorescence microscopy.
DNA and the proteins that interact with it form the foundation of life processes. These interactions determine whether certain genes are activated or suppressed, enable the recognition and repair of DNA damage, and ensure the transmission of genetic information. To better understand these dynamic processes and uncover their mechanisms, researchers need experimental approaches that enable continuous, real-time observation of individual proteins.
Why is it important to observe a single molecule?
According to Dr. M. Tutkus, the study’s co-author and principal investigator (PI), traditional biochemical methods often “hide” the behaviour of individual molecules because they measure the average value of a parameter across trillions of molecules.
“When we observe trillions of molecules at once, we see a parameter that reflects their overall average behavior. However, single-molecule methods allow us to see how a specific DNA–protein complex actually behaves – when it binds, how long it remains bound, or whether it moves along the DNA strand. This type of observation also allows us to exclude non-functional proteins that would otherwise distort the results,” the scientist explains.
The method used by VU LSC researchers is based on the so-called DNA flow-stretch system. Long DNA fragments are attached to a glass surface, labeled with DNA-intercalating fluorescent dyes, and stretched inside a specialized flow channel using liquid flow. As a result, the DNA molecules become effectively “visible” – they align linearly and can be observed using TIRF (total internal reflection fluorescence) microscopy.
According to Dr. M. Tutkus, stretching DNA in this way enables the simultaneous observation of individual molecules and their interactions.
“When a DNA molecule is stretched and positioned close to the surface, we can see with great precision what is happening at a specific location,” he says.
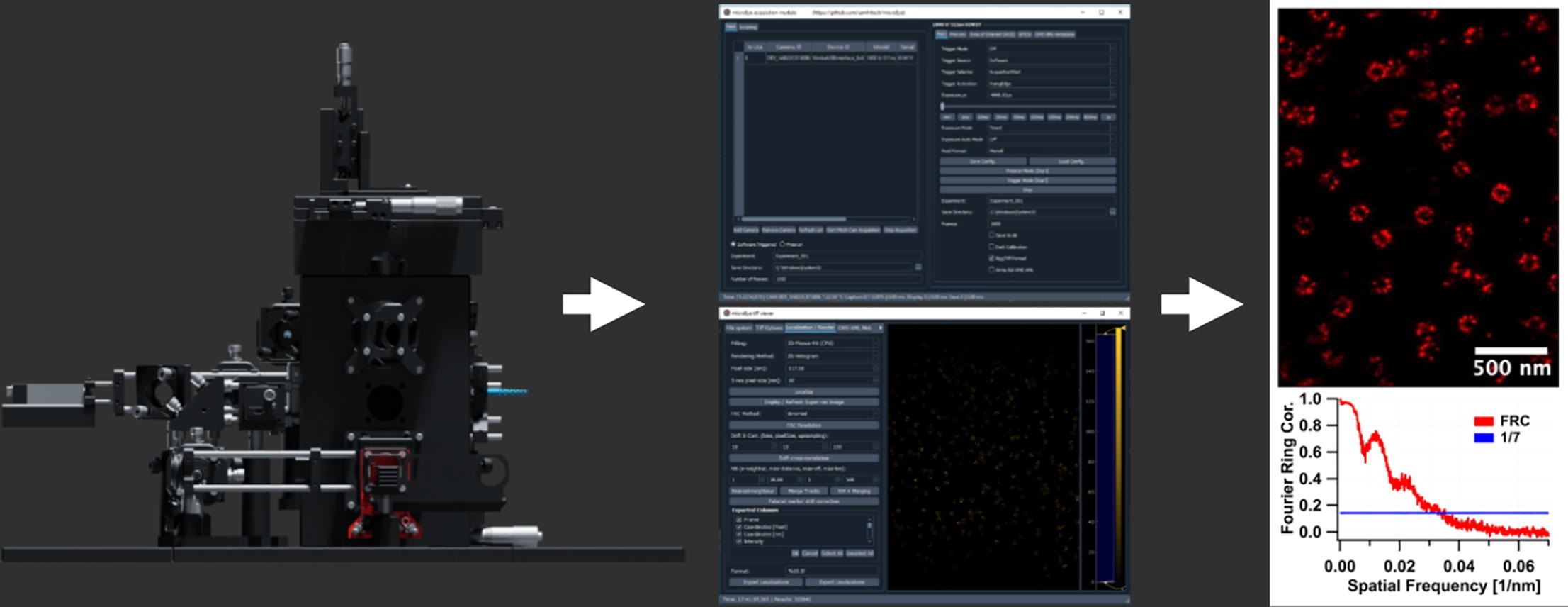 |
| The SR microscopy enables biomedical researchers to obtain structural and functional information on biomolecules that were inaccessible by any other method |
This advanced microscopy approach, developed by the research team, is essential for the method because it enables particularly clear visualization of labeled molecules.
From surface chemistry to real-time molecular observation
One of the most important aspects of the method is the careful preparation of the glass surface. The researchers optimized the surface-coating chemistry to keep DNA molecules firmly immobilized while minimizing nonspecific binding of DNA-interacting proteins to the glass surface.
The article describes all steps in detail – from preparing DNA fragments and functionalizing them with biotin or digoxigenin to attaching DNA to the surface at both ends and labeling these molecules with fluorescent dyes.
According to Dr. M. Tutkus, such a detailed description of the methodology is important so that it can be adopted by other laboratories.
“It was important for us not only to demonstrate that the method works but also to provide a clear protocol that other researchers could adopt. We have heard this repeatedly at various conferences. For many researchers, reliable and high-quality glass surface modification – which we have refined – is particularly important,” he says.
International collaboration between Lithuania and Taiwan
The chemical compounds required for glass surface modification were developed in collaboration with Prof. Chun-Jen Huang’s research team from Taiwan.
“This research group specializes in the synthesis of zwitterionic and other functionalized silatranes and is considered one of the global leaders in this field. The unique molecules they developed allow the formation of an extremely stable, thin, and uniform monolayer on the glass surface. Zwitterionic silatranes ensure efficient repulsion of nonspecific proteins, while amino-functionalized silatranes make it possible to attach DNA molecules to the surface,” the scientist explains.
It is worth noting that Chun-Jen Huang and Van Truck Vu are also co-authors of the article.
“Such international collaborations allow us to combine different areas of expertise. Surface chemistry is a critical element in experiments of this kind. The compounds developed by our partners enabled more stable and reliable DNA immobilization. We believe they will also be useful and relevant to other researchers,” says Dr. M. Tutkus.
A new method for simpler and more accessible studies in the future
The method has already been applied to study the interaction between DNA and proteins of the well-known CRISPR-Cas gene-editing systems, whose discovery was significantly advanced by the research team of Prof. Virginijus Šikšnys, a scientist at VU LSC.
According to Dr. M. Tutkus, methodological studies that introduce new research approaches can make such methods more accessible and easier to use. Over time, they may become widely adopted and turn into standard tools for developing and testing new DNA-interacting proteins.
The development of such new methods is important not only for fundamental molecular biology research but also for biotechnology applications in medicine.
“The better we understand molecular behaviour at the single-molecule level, the more precisely we will be able to control genetic processes and ultimately develop gene-editing systems that could one day be safely applied to treat human diseases,” says Dr. M. Tutkus.

