 |
Linas Mažutis, PhD Research Professor Group Leader Research ID: A-5408-2012 Email: Phone: +370 693 19333 |
Recent advances in high-throughput single technologies and computational methods have opened new horizons for biological and biomedical sciences. Just over the last few years, we have witnessed significant efforts to develop various analytical techniques to isolate, amplify and sequence the genetic material of individual cells. As the applications of single-cell sequencing continue to expand to all branches of life sciences there is a growing need for technological solutions that can deliver increased molecular sensitivity and reaction throughput at a reduced cost. Droplet microfluidics, a technology that enables pico- and nano-litre volume reactions, plays a major role in this endeavour. Our group are experts in droplet microfluidics technology for single-cell and many biological applications. Our group is pursuing research in cancer and immune system biology, aiming at better understanding of the genetic programs that drive tumour heterogeneity, progression and immune response.
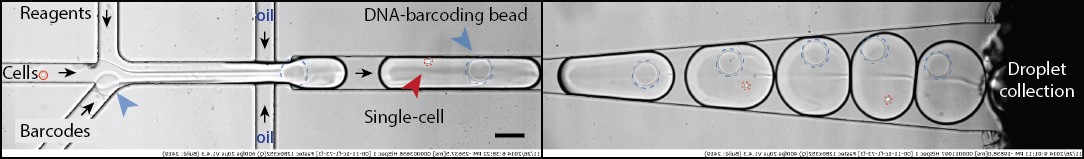
Fig 1. The principle of inDrops technique. Digital micrographs of cell encapsulation together with hydrogel beads and reagents. Scale bars, 100 μm. Cell loading into droplets with hydrogel beads and assay reagents occurs at the flow-focusing junction. Hydrogel bead ferries ssDNA primers attached to hydrogel polymer mesh via UV light-sensitive bond
In collaboration with Harvard University, our group has pioneered the droplet microfluidics technique inDrops (indexing Drops) for barcoding the transcriptome of individual cells (Klein, Cell, 2015). Since then, the technique has triggered immense attention among many scientists across different disciplines. We are applying inDrops and other techniques to better understand the gene expression programs that drive the development of complex diseases (e.g. tumours) and how the immune system responds. In collaboration with the Harvard Medical School (Dr A. Klein), we have studied the pluripotency of mouse embryonic cells [1], the T-cell activation in tumours [2] and the osteoblast role in lung adenocarcinoma [3], all single-cell level. In collaboration with Memorial Sloan Kettering Cancer Center and Columbia University (Dr. D. Pe’er), we have also shown that the T-cell exhibits a continuum of activated states to fight breast cancer [4] and, in a separate study, we have developed computational tools for recovering gene dropouts that are persistent in scRNA-Seq data [5].
SELECTED PUBLICATIONS
- Klein, M., Mazutis, L., Akartuna, I., Tallapragada, N., Veres, A., Li, V., Peshkin, L., Weitz, D. and Kirschner, M. Droplet barcoding for single cell transcriptomics applied to embryonic stem cells. Cell. 2015, 161(5): 1187–1201.
- Zemmour, D., Zilionis, R., Kiner, E., Klein, A. M., Mathis, D., Benoist, C. Single-cell gene expression reveals a landscape of regulatory T cell phenotypes shaped by the TCR. Nat Immunol. 2018, Mar; 19(3): 291–301.
- Engblom, C., Pfirschke, C., Zilionis, R., Da Silva Martins, J., Bos, S. A., Courties, G., Rickelt, S., Severe, N., Baryawno, N., Faget, J., Savova, V., Zemmour, D., Kline, J., Siwicki, M., Garris, C., Pucci, F., Liao, H. W., Lin, Y. J., Newton, A., Yaghi, O. K., Iwamoto, Y., Tricot, B., Wojtkiewicz, G. R., Nahrendorf, M., Cortez-Retamozo, V., Meylan, E., Hynes, R. O., Demay, M., Klein, A., Bredella, M. A., Scadden, D. T., Weissleder, R., Pittet, M. J. Osteoblasts remotely supply lung tumors with cancer-promoting SiglecFhigh neutrophils. Science. 2017, 12 01: 358(6367).
- Azizi, E., Carr, A., Plitas, G., Cornish, E. A., Konopacki, C., Prabhakaran, S., Nainys, J., Wu, K., Kiseliovas, V., Setty, M., Choi, K., Dao, P., Mazutis, L., Rudensky, Y. A. & Peer, D. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell. August 23, 2018, 174: 1–16.
- van Dijk, D., Sharma, R., Nainys, J., Yim, K., Kathail, P., Carr, A., Burdziak, C., Moon, R. K., Chaffer, C., Pattabiraman, D., Bierie, B., Mazutis, L., Wolf, G., Krishnaswamy, S., Peer, D. Recovering gene interactions from single-cell data using data diffusion. Cell. July 29, 2018, 174: 716–729.
Screening and Isolation of B-cells Producing Therapeutic Antibodies
Monoclonal antibodies constitute important tools for scientific research and are the basis of numerous successful therapeutics. However, traditional approaches to generate monoclonal antibodies against a desired target, such as hybridoma-based techniques and display library methods, are laborious and, due to fusion inefficiency and display bias, respectively, are unable to sample the entire antibody repertoire. We are working on a new platform to rapidly generate recombinant monoclonal antibodies. We use droplet microfluidics and a bead-based binding assay to directly identify and isolate individual rare cells that secrete target-binding antibody from a primary cell population with high throughput, screening more than one million cells per day. We then perform RT-PCR on individual sorted cells to recover the correctly paired heavy- and light-chain antibody sequences. We verify target-specific binding using ELISA. Our platform can facilitate rapid screening of an animal’s IgG-secreting cell repertoire to generate antigen specific recombinant antibodies and can also be adapted to isolate cells based on virtually any secreted product.
Transcriptional Profiling of Tumour
Tissue homeostasis is maintained by stem cells, whereas damaged tissues are repaired by facultative progenitors that are activated upon injury. The role of developmental plasticity in tumour progression and metastasis remains poorly understood and the extent to which tumour cells subvert regenerative processes during metastatic progression is unknown. In collaboration with biomedical and computational scientists, we applied inDrops technology to explore tumour cell heterogeneity through in lung cancer metastasis and to assess tumour cell plasticity. Using patient tumours as well as a mouse model of lung cancer metastasis, we identified the regenerative cell types and lineage promiscuity in untreated primary tumours and revealed a range of embryonic lung morphogenic states in metastases. We demonstrate an unexpected, developmental stage-specific differential sensitivity to natural killer cells that shapes the phenotypic landscape of latent metastasis-initiating cells.
Hydrogel Capsules for Single-Cell Multi-Step Processing
Droplet microfluidics technology provides a powerful approach to isolate and process millions of single cells simultaneously. However, multi-step reaction, including molecular biology and cell-based phenotypic screening assays, cannot be easily adapted to droplet format. To circumvent these limitations, we combined advantages offered by droplet-based and hydrogel-based systems to create capsules containing a thin, semi-permeable shell. The shell acts as a passive sieve retaining encapsulated, large molecular weight compounds while allowing smaller molecules (such as proteins) to diffuse through. We used an aqueous two-phase system (ATPS) composed of dextran and acrylate-modified polyethylene glycol to generate the biocompatible hydrogel particles and showcased a few examples of sequential reactions on encapsulated species. Specifically, we compared genome amplification reaction efficiency on Gram-negative and Gram-positive bacterial cells and found that the DNA amplification yields tend to be higher in the capsule-based system. Capsules readily sustained multiple pipetting steps when performing complex biochemical reactions, and, in contrast to solid-hydrogel beads, retained a significantly larger fraction of encapsulated bacteria.
